On December 30, 2019, a report indicated a cluster of patients with pneumonia of unknown etiology in Wuhan city, Hubei Province, China, was published on ProMED-mail. It was possibly related to contact with a local fish and wild animal market (Huanan Seafood Wholesale Market), where there was also sale of live animals. Most of the first reported patients visited the market about 1 month before onset. Deep sequencing analysis from lower respiratory tract samples indicated a novel coronavirus, which was named 2019 novel coronavirus (2019-nCoV) by WHO. In mid-January 2020, China began passenger transportation during or around the Chinese New Year which is equivalent to Christmas Holidays celebration in the West. A large number of people living in Wuhan population left the region by air carrier. Since January 17, the confirmed or suspected cases have dramatically increased. By 11:41 on February 05, 2020, China has reported 24363 confirmed cases and 23260 suspected cases including health-care workers, from all provinces, cities, autonomous regions and special administrative region (Hong Kong and Macao). Thailand, Japan, South Korea, and the USA also reported exported cases. There were 492 patients who died in China and 897 cases cured, at the time of this writing. The 2019-nCoV poses significant threats to international health.
Coronaviruses are enveloped non-segmented positive-sense RNA viruses belonging to the family Coronaviridae and the order Nidovirales and broadly distributed in humans and other mammals; The 2002–3 pandemic of severe acute respiratory syndrome (SARS) and the ongoing emergence of the Middle East respiratory syndrome coronavirus (MERS-CoV) demonstrate that coronaviruses (CoVs) are a significant public health threat The 2019-nCoV is a β CoV of group 2B with at least 70% similarity in genetic sequence to SARS-CoV. Different from both MERS-CoV and SARS-CoV, 2019-nCoV is the seventh member of the family of coronaviruses that infect humans. It may originate from Chinese horseshoe bats which are the natural reservoirs of (SARS-CoV) and can transmit from human to human.
The Shanghai Public Health Clinical Center is the designated hospital for diagnosis and management of infectious diseases and threats against public health in our country, and is also WHO designated training organization for new emerging infectious diseases. At the time of this writing, 56 confirmed patients with 2019-nCoV infection have been admitted to the Center.
The purpose of this study was to review the non-contrast chest CT findings in patients with positive new coronavirus nucleic acid antibody performed at the Center for Disease Control, Shangai, China.
On December 30, 2019, a report indicated a cluster of patients with pneumonia of unknown etiology in Wuhan city, Hubei Province, China, was published on ProMED-mail. It was possibly related to contact with a local fish and wild animal market (Huanan Seafood Wholesale Market), where there was also sale of live animals. Most of the first reported patients visited the market about 1 month before onset. Deep sequencing analysis from lower respiratory tract samples indicated a novel coronavirus, which was named 2019 novel coronavirus (2019-nCoV) by WHO. In mid-January 2020, China began passenger transportation during or around the Chinese New Year which is equivalent to Christmas Holidays celebration in the West. A large number of people living in Wuhan population left the region by air carrier. Since January 17, the confirmed or suspected cases have dramatically increased. By 11:41 on February 05, 2020, China has reported 24363 confirmed cases and 23260 suspected cases including health-care workers, from all provinces, cities, autonomous regions and special administrative region (Hong Kong and Macao). Thailand, Japan, South Korea, and the USA also reported exported cases. There were 492 patients who died in China and 897 cases cured, at the time of this writing. The 2019-nCoV poses significant threats to international health.
Infection
Human-to-human transmission of the virus has been confirmed[8] and occurs primarily via respiratory droplets from coughs and sneezes within a range of about 6 feet (1.8 m).[21][22] Viral RNA has also been found in stool samples from infected patients.[23]
It is possible that the virus can be infectious even during the incubation period, but this has not been proven,[24] and the WHO stated on 1 February 2020 that "transmission from asymptomatic cases is likely not a major driver of transmission" at this time.[25]
Human-to-human transmission of the virus has been confirmed[8] and occurs primarily via respiratory droplets from coughs and sneezes within a range of about 6 feet (1.8 m).[21][22] Viral RNA has also been found in stool samples from infected patients.[23]
It is possible that the virus can be infectious even during the incubation period, but this has not been proven,[24] and the WHO stated on 1 February 2020 that "transmission from asymptomatic cases is likely not a major driver of transmission" at this time.[25]
Reservoir
Animals sold for food were originally suspected to be the reservoir or intermediary hosts of SARS-CoV-2 because many of the first individuals found to be infected by the virus were workers at the Huanan Seafood Market.[26] A market selling live animals for food was also blamed in the SARS outbreak in 2003; such markets are considered to be incubators for novel pathogens.[27] The outbreak has prompted a temporary ban on the trade and consumption of wild animals in China.[28][29] However, some researchers have suggested that the Huanan Seafood Market may not be the original source of viral transmission to humans.[30][31]
Research into the origin of the 2003 SARS outbreak has resulted in the discovery of many SARS-like bat coronaviruses, most originating in the Rhinolophus genus of horseshoe bats. Two viral nucleic acid sequences from Rhinolophus sinicus published in 2015 and 2017 show a resemblance of 80% to SARS-CoV-2.[32][33][11] A third viral nucleic acid sequence from Rhinolophus affinis, "RaTG13" collected in Yunnan province, has a 96% resemblance to SARS-CoV-2.[9][34] For comparison, this amount of variation among viruses is similar to the amount of mutation observed over ten years in the H3N2 human influenza virus strain.[35]
Researchers from Guangzhou claim to have found a "99% identical" viral nucleic acid sequence in a pangolin sample.[36] As of 12 February 2020, the sequence remains unavailable, and all information comes from a university announcement.[37] Pangolins are protected under Chinese law, but poaching and trading of pangolins for traditional medicine remains common. A metagenomic study published in 2019 previously revealed that SARS-CoV was the most widely distributed coronavirus among a sample of Malayan pangolins.[38] Microbiologists and geneticists in Texas have independently found evidence of recombination in coronaviruses suggesting pangolin origins of SARS-CoV-2; they acknowledged remaining unknown factors while urging continued examination of other mammals.[39]
Animals sold for food were originally suspected to be the reservoir or intermediary hosts of SARS-CoV-2 because many of the first individuals found to be infected by the virus were workers at the Huanan Seafood Market.[26] A market selling live animals for food was also blamed in the SARS outbreak in 2003; such markets are considered to be incubators for novel pathogens.[27] The outbreak has prompted a temporary ban on the trade and consumption of wild animals in China.[28][29] However, some researchers have suggested that the Huanan Seafood Market may not be the original source of viral transmission to humans.[30][31]
Research into the origin of the 2003 SARS outbreak has resulted in the discovery of many SARS-like bat coronaviruses, most originating in the Rhinolophus genus of horseshoe bats. Two viral nucleic acid sequences from Rhinolophus sinicus published in 2015 and 2017 show a resemblance of 80% to SARS-CoV-2.[32][33][11] A third viral nucleic acid sequence from Rhinolophus affinis, "RaTG13" collected in Yunnan province, has a 96% resemblance to SARS-CoV-2.[9][34] For comparison, this amount of variation among viruses is similar to the amount of mutation observed over ten years in the H3N2 human influenza virus strain.[35]
Researchers from Guangzhou claim to have found a "99% identical" viral nucleic acid sequence in a pangolin sample.[36] As of 12 February 2020, the sequence remains unavailable, and all information comes from a university announcement.[37] Pangolins are protected under Chinese law, but poaching and trading of pangolins for traditional medicine remains common. A metagenomic study published in 2019 previously revealed that SARS-CoV was the most widely distributed coronavirus among a sample of Malayan pangolins.[38] Microbiologists and geneticists in Texas have independently found evidence of recombination in coronaviruses suggesting pangolin origins of SARS-CoV-2; they acknowledged remaining unknown factors while urging continued examination of other mammals.[39]
Phylogenetics and taxonomy
Genomic information
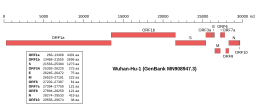 Genomic organisation of SARS-CoV-2
Genomic organisation of SARS-CoV-2
NCBI genome ID MN908947 Genome size 29,903 bases Year of completion 2020
SARS-CoV-2 belongs to the broad family of viruses known as coronaviruses. It is a positive-sense single-stranded RNA (+ssRNA) virus. Other coronaviruses are capable of causing illnesses ranging from the common cold to more severe diseases such as Middle East respiratory syndrome (MERS). It is the seventh known coronavirus to infect people, after 229E, NL63, OC43, HKU1, MERS-CoV, and the original SARS-CoV.[40]
Like the SARS-related coronavirus strain implicated in the 2003 SARS outbreak, SARS-CoV-2 is a member of the subgenus Sarbecovirus (Beta-CoV lineage B).[41][26][42] Its RNA sequence is approximately 30,000 bases in length.[7]
With a sufficient number of sequenced genomes, it is possible to reconstruct a phylogenetic tree of the mutation history of a family of viruses. By 12 January 2020, five genomes of SARS-CoV-2 had been isolated from Wuhan and reported by the Chinese Center for Disease Control and Prevention (CCDC) and other institutions;[7][43] the number of genomes increased to 81 by 11 February 2020.[44] A phylogenic analysis of the samples shows they are "highly related with at most seven mutations relative to a common ancestor", implying that the first human infection occurred in November or December 2019.[44]
On 11 February 2020, ICTV announced that according to existing rules that compute hierarchical relationships among coronaviruses on the basis of five conserved sequences of nucleic acids, the differences between what was then called 2019-nCoV and the virus strain from the 2003 SARS outbreak were insufficient to make it a separate viral species. Therefore, they identified 2019-nCoV as a strain of severe acute respiratory syndrome-related coronavirus.[1]

Genomic organisation of SARS-CoV-2
| |
| NCBI genome ID | MN908947 |
|---|---|
| Genome size | 29,903 bases |
| Year of completion | 2020 |
SARS-CoV-2 belongs to the broad family of viruses known as coronaviruses. It is a positive-sense single-stranded RNA (+ssRNA) virus. Other coronaviruses are capable of causing illnesses ranging from the common cold to more severe diseases such as Middle East respiratory syndrome (MERS). It is the seventh known coronavirus to infect people, after 229E, NL63, OC43, HKU1, MERS-CoV, and the original SARS-CoV.[40]
Like the SARS-related coronavirus strain implicated in the 2003 SARS outbreak, SARS-CoV-2 is a member of the subgenus Sarbecovirus (Beta-CoV lineage B).[41][26][42] Its RNA sequence is approximately 30,000 bases in length.[7]
With a sufficient number of sequenced genomes, it is possible to reconstruct a phylogenetic tree of the mutation history of a family of viruses. By 12 January 2020, five genomes of SARS-CoV-2 had been isolated from Wuhan and reported by the Chinese Center for Disease Control and Prevention (CCDC) and other institutions;[7][43] the number of genomes increased to 81 by 11 February 2020.[44] A phylogenic analysis of the samples shows they are "highly related with at most seven mutations relative to a common ancestor", implying that the first human infection occurred in November or December 2019.[44]
On 11 February 2020, ICTV announced that according to existing rules that compute hierarchical relationships among coronaviruses on the basis of five conserved sequences of nucleic acids, the differences between what was then called 2019-nCoV and the virus strain from the 2003 SARS outbreak were insufficient to make it a separate viral species. Therefore, they identified 2019-nCoV as a strain of severe acute respiratory syndrome-related coronavirus.[1]
Structural biology
Protein modeling experiments on the spike (S) protein of the virus suggest that it has sufficient affinity to the angiotensin converting enzyme 2 (ACE2) receptors of human cells to use them as a mechanism of cell entry.[45] On 22 January 2020, a group in China working with the full virus genome and a group in the United States using reverse genetics methods independently and experimentally demonstrated that ACE2 could act as the receptor for SARS-CoV-2.[46][47][48][49] Studies have shown that SARS-CoV-2 has a higher affinity to human ACE2 than the original SARS virus strain.[50] An atomic-level image of the S protein has been created using cryogenic electron microscopy.[51][52]
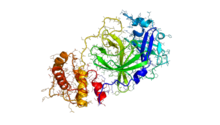
To look for potential protease inhibitors, the viral 3C-like protease M(pro) from the ORF1a polyprotein has also been modeled for drug docking experiments. Innophore has produced two computational models based on SARS protease,[53] and the Chinese Academy of Sciences has produced an unpublished experimental structure of a recombinant 2019-nCoV protease.[54] In addition, researchers have modeled the structures of all mature peptides in the SARS-CoV-2 genome using I-TASSER and Swiss-model.[55][56]
Digitally colorized electron micrographs of SARS-CoV-2
Protein modeling experiments on the spike (S) protein of the virus suggest that it has sufficient affinity to the angiotensin converting enzyme 2 (ACE2) receptors of human cells to use them as a mechanism of cell entry.[45] On 22 January 2020, a group in China working with the full virus genome and a group in the United States using reverse genetics methods independently and experimentally demonstrated that ACE2 could act as the receptor for SARS-CoV-2.[46][47][48][49] Studies have shown that SARS-CoV-2 has a higher affinity to human ACE2 than the original SARS virus strain.[50] An atomic-level image of the S protein has been created using cryogenic electron microscopy.[51][52]
To look for potential protease inhibitors, the viral 3C-like protease M(pro) from the ORF1a polyprotein has also been modeled for drug docking experiments. Innophore has produced two computational models based on SARS protease,[53] and the Chinese Academy of Sciences has produced an unpublished experimental structure of a recombinant 2019-nCoV protease.[54] In addition, researchers have modeled the structures of all mature peptides in the SARS-CoV-2 genome using I-TASSER and Swiss-model.[55][56]
Digitally colorized electron micrographs of SARS-CoV-2
Epidemiology
The first known human infection by the strain occurred in early December 2019.[57][30] Proliferation of SARS-CoV-2 was first detected in Wuhan, China, in mid-December 2019, likely originating from a single infected animal.[30] The virus subsequently spread to all provinces of China and to more than two dozen other countries in Asia, Europe, North America, Africa, and Oceania.[58] Human-to-human spread of the virus has been confirmed in all of these regions[8][59][60][61] except Africa.[62] On 30 January 2020, SARS-CoV-2 was designated a Public Health Emergency of International Concern by the WHO.[14][63]
As of 20 February 2020 (15:30 UTC), there were 77,923 confirmed cases of infection, of which 76,291 were within mainland China.[58] One mathematical model estimated the number of people infected in Wuhan alone at 75,815 as of 25 January 2020.[64] Nearly all cases outside China have occurred in people who either traveled from Wuhan, or were in direct contact with someone who traveled from the area.[65][66] While the proportion of infections that result in confirmed infection or progress to diagnosable disease remains unclear,[67][68] the total number of deaths attributed to the virus was 2,362 as of 22 February 2020 (15:15 UTC); over 95% of all deaths have occurred in Hubei province,[58] where Wuhan is located.
The basic reproduction number (, pronounced R-nought or R-zero)[69] of the virus has been estimated to be between 1.4 and 3.9.[70][71][72][73] This means that, when unchecked, the virus typically results in 1.4 to 3.9 new cases per established infection. It has been established that the virus is able to transmit along a chain of at least four people.[74] As of 19 February 2020, infection fatality ratio (IFR) estimates range from 0.3% to 1%, and the confirmed case fatality rate (CFR) within China is 2.3%.[75]
source: en.wikipedia.org/wiki/Severe_acute_respiratory_syndrome_coronavirus_2
The first known human infection by the strain occurred in early December 2019.[57][30] Proliferation of SARS-CoV-2 was first detected in Wuhan, China, in mid-December 2019, likely originating from a single infected animal.[30] The virus subsequently spread to all provinces of China and to more than two dozen other countries in Asia, Europe, North America, Africa, and Oceania.[58] Human-to-human spread of the virus has been confirmed in all of these regions[8][59][60][61] except Africa.[62] On 30 January 2020, SARS-CoV-2 was designated a Public Health Emergency of International Concern by the WHO.[14][63]
As of 20 February 2020 (15:30 UTC), there were 77,923 confirmed cases of infection, of which 76,291 were within mainland China.[58] One mathematical model estimated the number of people infected in Wuhan alone at 75,815 as of 25 January 2020.[64] Nearly all cases outside China have occurred in people who either traveled from Wuhan, or were in direct contact with someone who traveled from the area.[65][66] While the proportion of infections that result in confirmed infection or progress to diagnosable disease remains unclear,[67][68] the total number of deaths attributed to the virus was 2,362 as of 22 February 2020 (15:15 UTC); over 95% of all deaths have occurred in Hubei province,[58] where Wuhan is located.
The basic reproduction number (, pronounced R-nought or R-zero)[69] of the virus has been estimated to be between 1.4 and 3.9.[70][71][72][73] This means that, when unchecked, the virus typically results in 1.4 to 3.9 new cases per established infection. It has been established that the virus is able to transmit along a chain of at least four people.[74] As of 19 February 2020, infection fatality ratio (IFR) estimates range from 0.3% to 1%, and the confirmed case fatality rate (CFR) within China is 2.3%.[75]
source: en.wikipedia.org/wiki/Severe_acute_respiratory_syndrome_coronavirus_2






Comments
Post a Comment